EME: GDS
To simulate more complex structures, a GDS import function is used to support any geometry that can be created on a mask GDS file. This example shows how an s-bend structure can be created in a GDS file, imported to EMode, and simulated as part of a directional coupler.
This code example is licensed under the BSD 3-Clause License.
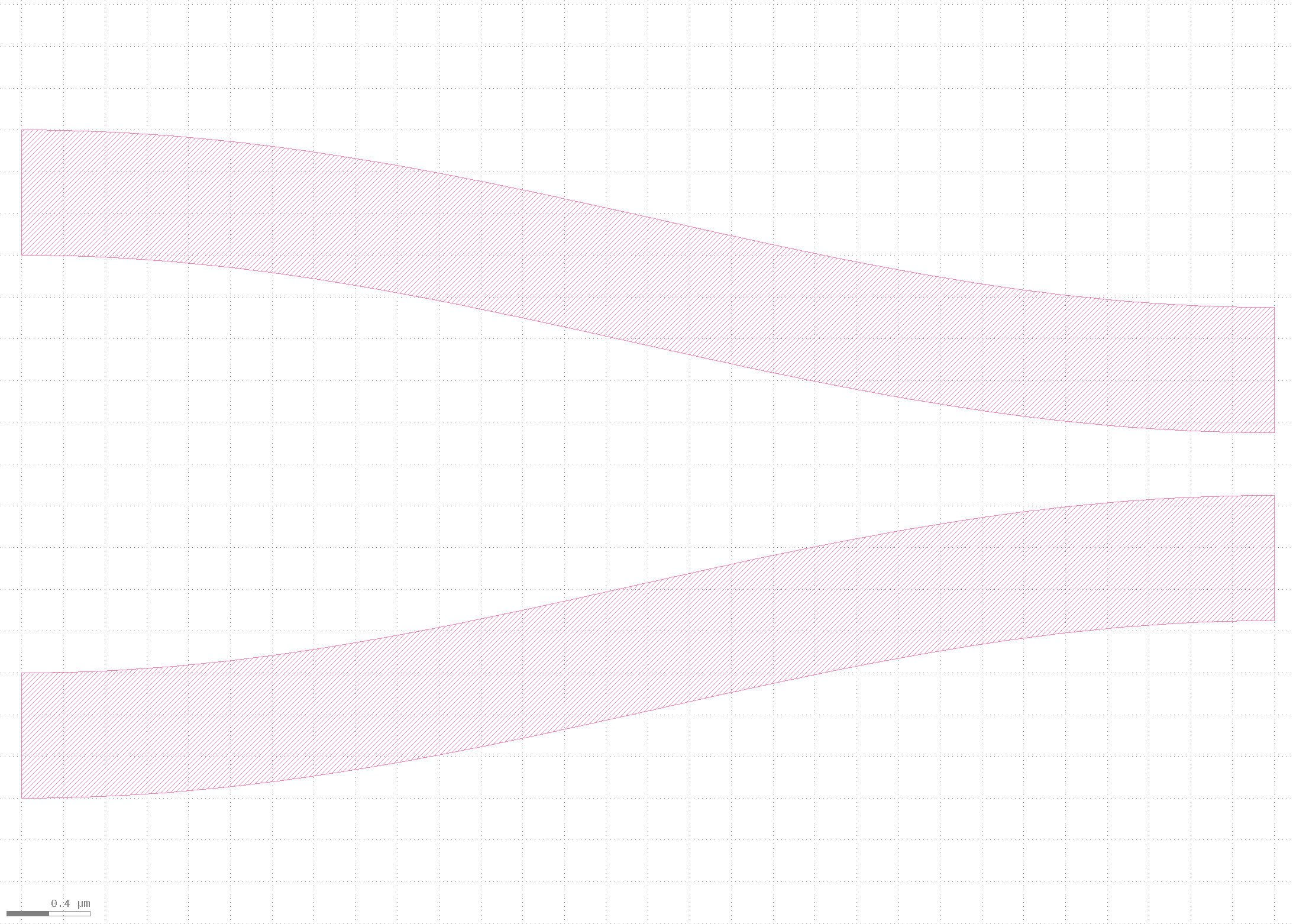
First, the GDS is created.
from phidl import CrossSection, Device
import phidl.path as pp
from phidl import quickplot as qp
## Parameters
width_wg = 0.6 # [um]
gap_input = 2.0 # [um]
gap_output = 0.3 # [um]
L_bend = 6.0 # [um]
## Create the first CrossSection
X1 = CrossSection()
X1.add(width = width_wg,
offset = -gap_input/2 - width_wg/2, layer = 1,
name = 'wg1', ports = ('in1', 'out1'))
X1.add(width = width_wg,
offset = gap_input/2 + width_wg/2, layer = 1,
name = 'wg2')
## Create the second CrossSection
X2 = CrossSection()
X2.add(width = width_wg,
offset = -gap_output/2 - width_wg/2, layer = 1,
name = 'wg1', ports = ('in2', 'out2'))
X2.add(width = width_wg,
offset = gap_output/2 + width_wg/2, layer = 1,
name = 'wg2')
## Transition from first to second CrossSection
Xtrans = pp.transition(cross_section1 = X1, cross_section2 = X2, width_type = 'sine')
P3 = pp.straight(length = L_bend)
WG_trans = P3.extrude(Xtrans)
## Place all CrossSection extrusions
D = Device()
wgt = D << WG_trans
## Quickplot to view
qp(D)
## Save gds file
D.write_gds('sbend.gds')

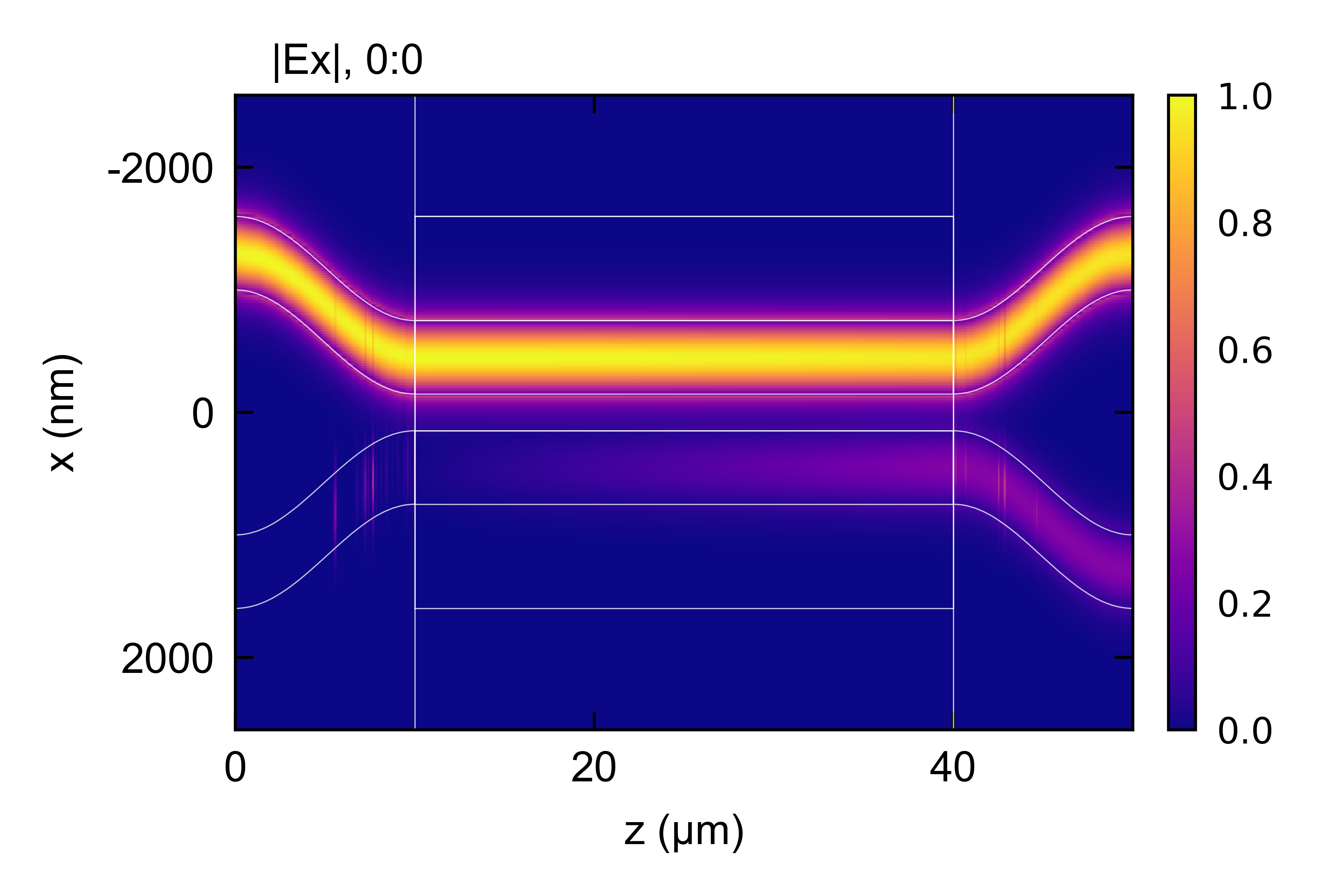
Next, the directional coupler device is created and simulated for a short coupler length. The sweep function is used to optimize the coupler length for 100 % coupling, and the propagation is re-plotted.
import emodeconnection as emc
from matplotlib import pyplot as plt
import numpy as np
## Set simulation parameters
wavelength = 1550 # [nm] wavelength
dx, dy = 20, 20 # [nm] resolution
h_core = 220 # [nm] waveguide core height
h_clad = 1200 # [nm] waveguide top and bottom clad
width_wg = 600 # [nm]
gap = 300 # [nm]
input_gap = 2000 # [nm]
window_height = h_core + h_clad*2
num_modes = 2 # [-] number of modes
BC = 'TE-v'
## Connect and initialize EMode
em = emc.EMode()
## Settings
em.settings(
wavelength = wavelength, x_resolution = dx, y_resolution = dy,
window_height = window_height, num_modes = num_modes,
boundary_condition = BC, background_material = 'SiO2')
## Draw shapes
em.shape(name = 'BOX', material = 'SiO2',
height = h_clad)
em.shape(name = 'core', material = 'Si',
height = h_core, etch_depth = h_core)
## Create profiles
em.settings(window_width = 6200)
em.label_profile(name = 'slab')
em.settings(window_width = 2600)
em.shape(name = 'core', mask = width_wg)
em.label_profile(name = 'io_waveguide')
em.settings(window_width = 2600)
em.shape(name = 'core', mask = [width_wg, width_wg],
mask_offset = [-gap/2-width_wg/2, gap/2+width_wg/2])
em.label_profile(name = 'coupler')
## Draw EME sections
em.section(name = 'input', section_type = 'straight',
profile = 'io_waveguide', length = 1000,
offset = input_gap/2 + width_wg/2)
em.section(name = 'sbend', section_type = 'gds',
gds = 'sbend.gds', profile = 'slab',
shape_to_mask = 'core',
length = 10e3) # modify length of gds
em.section(name = 'coupler', section_type = 'straight',
profile = 'coupler', length = 30e3)
em.section(section_type = 'copy',
section_name = 'sbend', mirror = True)
em.section(name = 'output', section_type = 'straight',
profile = 'io_waveguide', length = 1000,
offset = -input_gap/2 - width_wg/2)
## Run EME and plot results
em.EME()
em.plot()
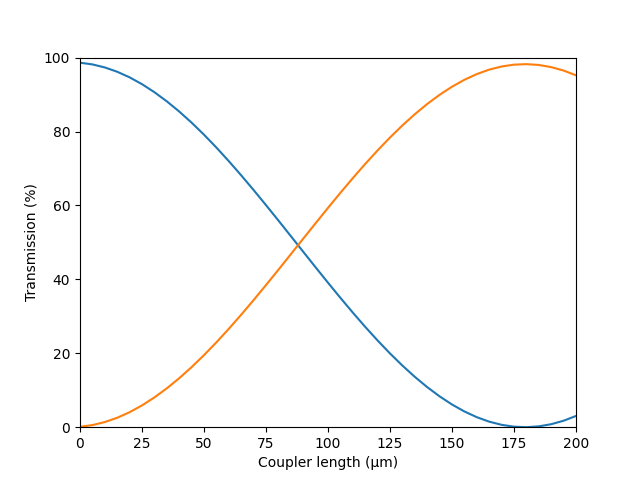
## Sweep the coupler length and plot results
data = em.sweep(key = 'section, coupler, length',
values = np.arange(0, 200.1e3, 10000),
result = ['S_matrix'])
S = data['S_matrix']
plt.plot(data['values']*1e-3, np.abs(S[:,1,0,0])**2*100)
plt.xlabel("Coupler length (\u03BCm)")
plt.ylabel("Transmission (%)")
plt.autoscale(enable = True, axis = 'x', tight = True)
plt.ylim([0,100])
plt.savefig('transmission_plot.png', dpi = 300, bbox_inches = 'tight')
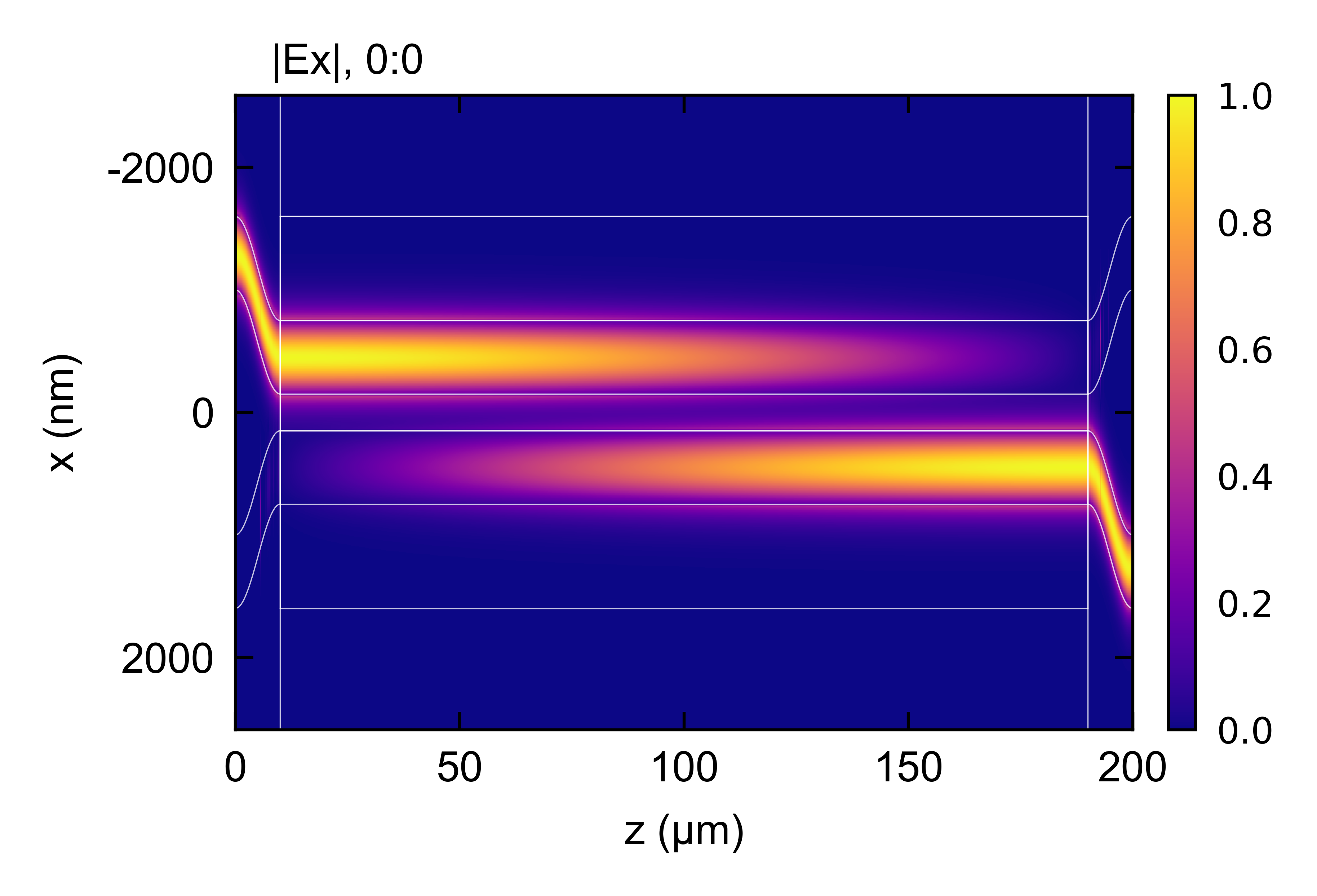
## Plot final design for 100 % coupler
em.section(name = 'coupler', length = 180e3)
em.EME()
em.plot()
## Close EMode
em.close()
Console output:
EMode3D 0.2.6 - email
Meshing completed in 0.3 sec
Meshing completed in 0.1 sec
Meshing completed in 0.1 sec
Solving S-matrices...
Solving section: input...Meshing completed in 0.1 sec
completed in 1.5 sec
Solving section: sbend...
Meshing completed in 0.1 sec
completed in 31.1 sec
Solving section: coupler... completed in 0.2 sec
Solving section: 3...
completed in 4.0 sec
Solving section: output... completed in 0.1 sec
completed in 0.1 sec
Meshing completed in 0.2 sec
Sweeping section parameter 'length'...
Solving EME: length = 200000.0... completed in 1.1 sec
Solving EME: length = 190000.0... completed in 1.1 sec
Solving EME: length = 180000.0... completed in 1.2 sec
Solving EME: length = 170000.0... completed in 1.2 sec
Solving EME: length = 160000.0... completed in 1.2 sec
Solving EME: length = 150000.0... completed in 1.2 sec
Solving EME: length = 140000.0... completed in 1.5 sec
Solving EME: length = 130000.0... completed in 1.5 sec
Solving EME: length = 120000.0... completed in 1.6 sec
Solving EME: length = 110000.0... completed in 1.4 sec
Solving EME: length = 100000.0... completed in 1.6 sec
Solving EME: length = 90000.0... completed in 1.3 sec
Solving EME: length = 80000.0... completed in 1.3 sec
Solving EME: length = 70000.0... completed in 1.3 sec
Solving EME: length = 60000.0... completed in 1.4 sec
Solving EME: length = 50000.0... completed in 1.3 sec
Solving EME: length = 40000.0... completed in 1.5 sec
Solving EME: length = 30000.0... completed in 1.6 sec
Solving EME: length = 20000.0... completed in 1.5 sec
Solving EME: length = 10000.0... completed in 1.5 sec
Solving EME: length = 0.0... completed in 1.8 sec
completed in 29.0 sec
Solving S-matrices...
Solving section: input... completed in 0.4 sec
Solving section: sbend... completed in 0.4 sec
Solving section: coupler... completed in 0.4 sec
Solving section: 3... completed in 0.3 sec
Solving section: output... completed in 0.1 sec
completed in 0.1 sec
Exited EMode
Figures: